A twinkle of light in a microscope’s eye
Researchers are using biological glow sticks and finely-tuned cameras to look deeper within cells.
Marissa Shieh • December 21, 2016
It’s a crowded party in the body, and biologists want to see what’s going on. New fluorescence microscopy techniques build on the ones used to capture this image of neurons. These new techniques may shed fresh light on proteins. [Image credit: Jan Schmoranzer from the 2011 Olympus BioScapes Digital Imaging Competition | CC BY 3.0]
Fireflies are easy to find at night — but not in the day. Their tails glow bright in the darkness. Yet, in sunshine, it’s difficult to compete with everything else. By using the same concept of pinpointing objects in the darkness with emitted light, fluorescence microscopy has helped scientists look inside cells for more than a century.
Unfortunately, that technology had a limit. Researchers could not look deep into proteins to see how the tiny molecular machines moved and interacted in living cells. Since proteins do many of a cell’s most crucial jobs — including fighting infections, carrying messages and initiating chemical reactions — poor image resolution has hindered the search for better medicines and keener insight into aging.
Under a conventional fluorescence microscope, proteins look like blurs of light instead of pinpricks, as if those fireflies were floating in a fog, grouped too closely together. But now that view may be sharpening.
In a paper published in July 2016 in Nature Nanotechnology, researchers associated with Harvard University’s Wyss Institute for Biologically Inspired Engineering revealed their updated toolkit, called discrete molecular imaging, which can clearly locate and count individual sections of proteins just five nanometers apart, a distance 20,000 times thinner than a sheet of paper. Previous techniques could only distinguish between protein sections 10 to 20 nanometers apart.
The improved resolution “is a game-changer,” says Katharina Gaus of the University of New South Wales, who is using the DNA-PAINT tool to learn how T cells in our immune system communicate with one another.
DNA-PAINT uses complimentary strands of DNA that light up when joined. One strand binds to the protein of interest, and the other drifts through the medium until it encounters the first strand. When they meet, the fluorescent tag on the free-floating strand lights up.
Looking through a microscope, the scientist sees this effect as pinpricks of light, then darkness as the strands separate, and then light once more, as the unattached strands play musical chairs.
Above is an animation of DNA-PAINT at work. By binding and separating fragments, scientists can produce pinpoints of light to help illuminate our understanding of proteins. [Video credit: Wyss Institute for Biologically Inspired Engineering at Harvard University]
As part of the larger, updated toolkit, a camera records the blinking lights and a computer compiles the images to form a complete picture of the protein.
It’s difficult to avoid blurring the composite image at this scale, so the Harvard team wrote an algorithm correcting the drift of the instruments to within one nanometer of accuracy.
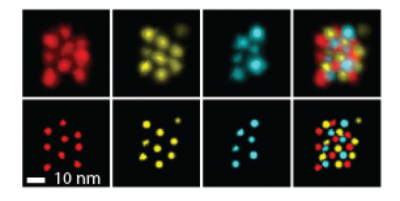

Each color corresponds to a different subsection of a protein. The lower set of images has been corrected by the computer algorithm. [Photo credit: Wyss Institute for Biologically Inspired Engineering]
Biologists hope to use these new tools to locate the positions of amino acids in proteins and investigate age-old questions about our cells: how proteins fold, what enzymes look like and how the proteins embedded in cell membranes work. Previous technology required tiny, frozen, or crystallized samples — a set-up divorced from how proteins naturally exist in our bodies. But the new techniques will allow biologists to use live tissue and cells.
When fully developed, these imaging techniques “could potentially improve all the ways people [analyze biological] samples,” suggests Mingjie Dai, the postdoctoral researcher on the Harvard team.
As for the overall promise of the technology, Dai says, “there is no hard limit.”